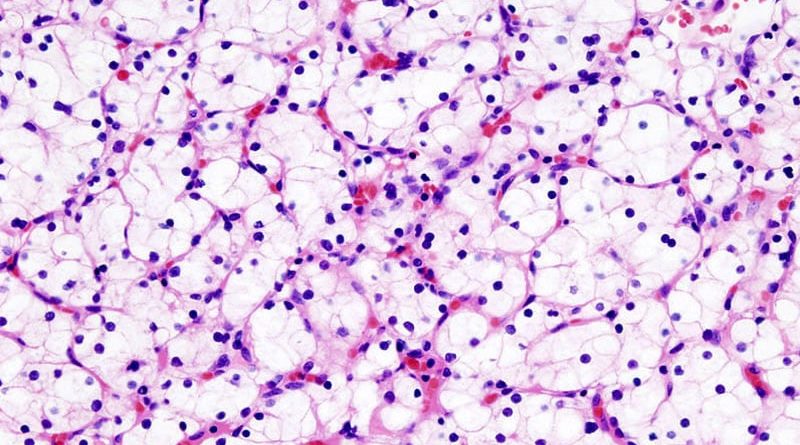
Signature Predicts Metastatic Survival in Clear Cell RCC
In this preclinical study, a single-transcriptome analysis revealed a distinct transcriptional signature that was predictive of metastatic potential and patient survival for patients with clear cell renal cell carcinoma (ccRCC). The analysis characterized the tumor microenvironment in ccRCC so as to identify therapeutic targets and to further biologic understanding of the disease. The study covered in this summary was published on bioRxiv.org as a preprint and has not yet been peer reviewed.
Key Takeaways
-
For patients with ccRCC, single-cell transcriptomic analysis of tumor cells from the primary tumors revealed a distinct transcriptional signature that can potentially be used to predict metastatic potential and patient survival.
-
Stromal cell analysis within the tumor microenvironment demonstrated vascular remodeling within the endothelial cells and fibroblast proliferation associated with poor survival.
-
A cell-to-cell interaction analysis, produced by computer modeling, highlighted the CXCL9/CXCL10-CXCR3 axis and the CD70-CD27 axis as potential therapeutic targets.
Why This Matters
-
The study contributes to the understanding that is required to improve therapies for metastatic ccRCC by determining how tumor cells are interacting with their microenvironment.
-
The study helps to elucidate the specific relationship between stromal cells and tumor cells within the tumor microenvironment to advance understanding of carcinogenesis and cancer progression.
-
Dissecting the cellular and molecular landscape of ccRCC can facilitate new avenues of therapeutic intervention and ultimately result in better treatments for patients suffering from ccRCC.
-
The study provides a valuable resource for further explorations into critical cellular relationships and signaling axes that remodel the tumor microenvironment, resulting in a tumor-suppressive environment.
Study Design
-
The study analyzed fresh patient samples of ccRCC tumors and compared treatment-naive primary tumor tissue to matched adjacent normal kidney tissue as well as samples collected from patients with bone metastases.
-
Nine patients were diagnosed initially with ccRCC. Two of those patients had clinical metastases at multiple sites at the time of diagnosis. In total, 157,881 cells were sequenced (ccRCC: 122,054 cells; normal kidney: 35,827 cells). Samples were integrated using a joint analysis of the heterogeneous samples.
-
Tumor tissues and adjacent normal kidney tissue collected from the same patient permitted a matched comparison and helped to control for interindividual variation.
-
Single-cell transcriptomic analysis of primary tumor cells was conducted.
Key Results
-
Tumor cells are transcriptionally similar to a subset of proximal tubule cells, which may be an indication of the tumor cell of origin.
-
The ccRCC cells are highly vascularized tumors with disorganized vasculature, endothelial cells, and pericytes.
-
The synchronous comparison of primary tumor and bone-metastatic tumor tissues from two patients who presented with de novo metastases revealed a specific metastatic signature associated with poor prognosis.
-
The stromal cells within RCC tumors show the highest transcriptional difference of analyzed cell types in comparison to adjacent normal kidney.
-
Exhausted T cells and immunosuppressive cell populations, including Tregs and Macro-2 cells, were found in the complex immune microenvironment of ccRCC.
-
Ligand-receptor cell interaction analysis highlighted the CXCL9/CXCL10-CXCR3 axis and the CD70-CD27 axis as potential therapeutic targets.
Limitations
-
Study limitations were not discussed by the authors.
Disclosures
-
The study received no commercial funding.
-
One author serves on the Scientific Advisory Board to Celsius Therapeutics Inc and Biomage Inc. One author is director and a shareholder for Agios Therapeutics and Editas Medicines; is a founder, director, and scientific advisory board member for Magenta Therapeutics and LifeVault Bio; and is a founder of Clear Creek Bio. One author is a founder and shareholder of Clear Creek Bio.
This is a summary of a preprint research study, “A New Transcriptional Metastatic Signature Predicts Survival in Clear Cell Renal Cell Carcinoma,” published July 1 on bioRxiv.org and led by Peter V. Kharchenko of Harvard Medical School, Boston. This study has not yet been peer reviewed. The full text of the study can be found on bioRxiv.org.
For more news, follow Medscape on Facebook, Twitter, Instagram, and YouTube.
Source: Read Full Article