Exploring how fungal infections spread in the human lung
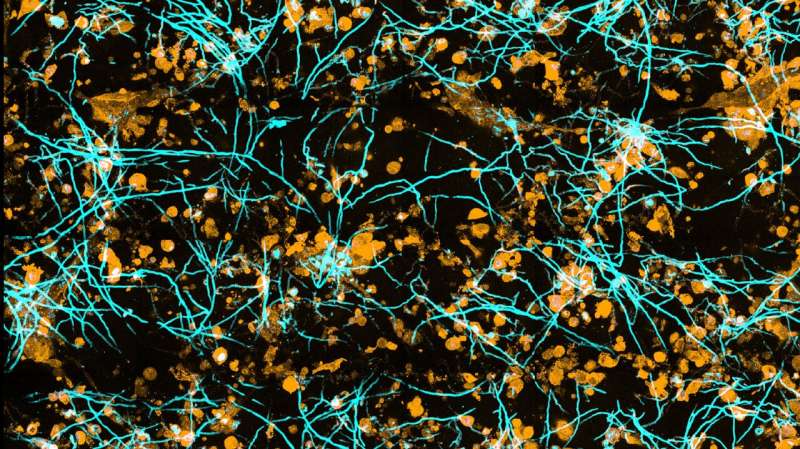
A chip-based infection model developed by researchers in Jena, Germany, enables live microscopic observation of damage to lung tissue caused by the invasive fungal infection aspergillosis. The team developed algorithms to track the spread of fungal hyphae as well as the response of immune cells. The development is based on a “lung-on-chip” model also developed in Jena and can help reduce the number of animal experiments. The results were presented in the journal Biomaterials.
Aspergillosis is a mold infection caused by Aspergillus fumigatus, which often affects the lungs. The disease can be fatal, especially in immunocompromised individuals. In these cases, invasive aspergillosis usually occurs with fungal hyphae invading blood vessels. So far, there are only a few active substances that can combat such fungal infections. “That’s why it was so important for us to be able to represent this invasive growth in a model,” says Marie von Lilienfeld-Toal, who co-led the study. The internist is a professor at the Department of Internal Medicine II at Jena University Hospital and conducts research at the Leibniz Institute for Natural Product Research and Infection Biology—Hans Knöll Institute (Leibniz-HKI) in Jena, Germany.
The new aspergillosis infection model should help to better observe both the growth of the fungus and the reaction of the immune system and to find possible new approaches for therapies. In addition, new active substances can be tested. The expertise for this is available in Jena: Organ chips have long been developed at the university hospital. The startup Dynamic42, which manufactures the lung chips used in the study, was founded there. First author Mai Hoang also joined the company after completing her doctorate.

From organ model to infection model
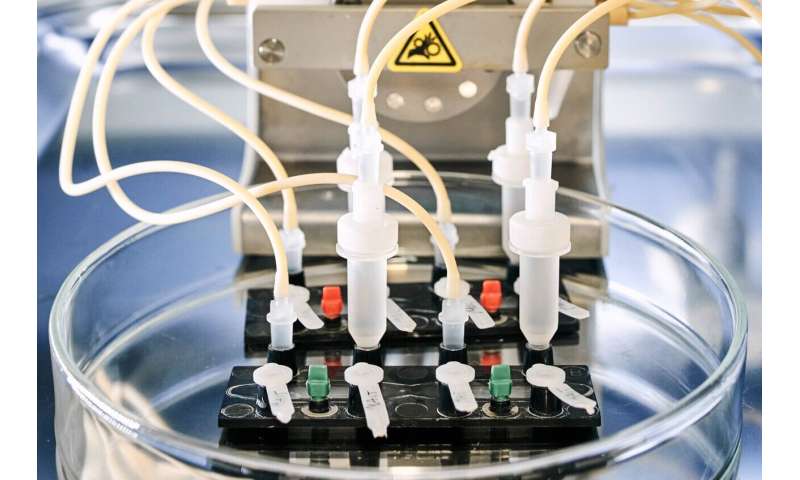
“With the help of the chip, we can observe and quantify aspergillosis in 3D live, under the microscope,” says study leader Marc Thilo Figge. He is head of the Applied Systems Biology research group at Leibniz-HKI and a professor at Friedrich Schiller University in Jena. The organ model consists of two layers of cells separated by an artificial membrane. One layer is exposed to air and consists of surface cells of the lung. The other layer consists of blood vessel cells, with a blood-like nutrient solution continuously flowing past them.
To this model, the researchers then added the fungus. “In this way, we turned the organ model into an infection model,” explains Susann Hartung, a member of the Infections in Hematology/Oncology group at Leibniz-HKI and one of the three first authors. The difficulty, she says, was establishing the right severity of infection. “If we add too much Aspergillus fumigatus to the model, the lung cells die. If there is too little, we don’t see anything,” the molecular biologist adds.
Human immune cells or various drugs, for example, can then be added to this system, as the research team shows in the current study.
Algorithm for image evaluation
Evaluating the three-dimensional microscopic data presented a major challenge. “If we just look at the images, we get a sense of the progression of the infection, but we can’t quantify it. For that, we need algorithms that can distinguish fungal hyphae or immune cells from tissue cells as well as the surrounding environment,” explains Zoltán Cseresnyés, who is also a first author of the paper. He is a member of Figge’s team who is specialized in automated image analysis.
To make it possible for the computer to distinguish between them, the different cell types are color-coded using fluorescent dyes. “For example, the intensity of the fluorescence can be used to determine how many fungi an immune cell has eaten,” Cseresnyés explains.
Source: Read Full Article